Check the quality of the anatomical coregistration
The following code makes a number of figures that can be used as quality control for the procedure to coregister the MRI with the MEG.
Please note that this is an example where the coregistration was not completely correct. The head is tilted to the left.
%% load the required data
load headmodel_mri.mat
load headshapeMEG.mat % from the fiff file
load mri_resliced.mat % resliced
load mri_segmented.mat
load sens.mat
%% figure 1
figure
ft_plot_sens(sens, 'unit', 'mm')
ft_plot_headshape(headshapeMEG, 'unit', 'mm')
ft_plot_headmodel(headmodel_mri, 'unit', 'mm')
ft_plot_axes([], 'unit', 'mm');
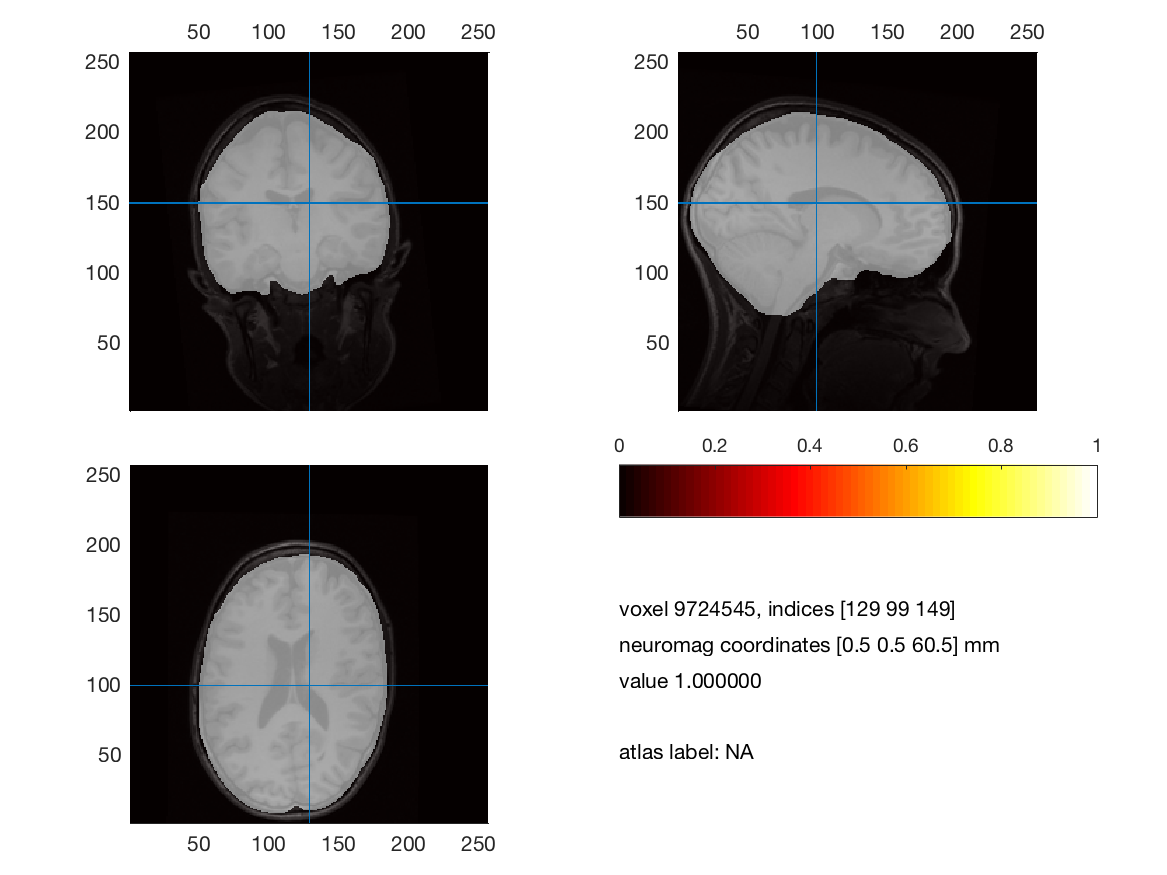
%% figure 2, MRI anatomy and brain segmentation
cfg = [];
cfg.anaparameter = 'anatomy';
cfg.funparameter = 'brain';
cfg.location = [0 0 60];
ft_sourceplot(cfg, mri_segmented)
%% figure 3 and 4, MRI anatomy and headmodel
location = [0 0 60];
figure
ft_plot_ortho(mri_resliced.anatomy, 'transform', mri_resliced.transform, 'location', location, 'intersectmesh', headmodel_mri.bnd)
figure
ft_plot_montage(mri_resliced.anatomy, 'transform', mri_resliced.transform, 'intersectmesh', headmodel_mri.bnd)
%% figure 5, MRI scalp surface and polhemus headshape
cfg = [];
cfg.tissue = 'scalp';
cfg.method = 'isosurface';
cfg.numvertices = 10000;
scalp = ft_prepare_mesh(cfg, mri_segmented);
figure
ft_plot_mesh(scalp, 'facecolor', 'skin')
lighting phong
camlight left
camlight right
material dull
alpha 0.5
ft_plot_headshape(headshapeMEG, 'vertexcolor', 'k');
%% figure 6, MRI and anatomical landmarks
figure
for i=1:3
subplot(2,2,i)
title(headshapeMEG.fid.label{i});
location = headshapeMEG.fid.pos(i,:);
ft_plot_ortho(mri_resliced.anatomy, 'transform', mri_resliced.transform, 'style', 'intersect', 'location', location, 'plotmarker', location, 'markersize', 5, 'markercolor', 'y')
end
%% figure 7, MRI scalp surface and anatomical landmarks
figure
ft_plot_mesh(scalp, 'facecolor', 'skin')
lighting phong
camlight left
camlight right
material dull
alpha 0.3
ft_plot_mesh(headshapeMEG.fid, 'vertexcolor', 'k', 'vertexsize', 10);