How to change the MRI orientation, the voxel size or the field-of-view?
The function ft_volumereslice allows you to
- align the anatomical MRI or functional volume along the x,y,z axis, i.e. put it right side up
- ensure that the voxels are isotropic, i.e. having equal size in all 3 dimensions
- zoom in or on a part of the volume, i.e. changing the number of voxels and FOV
- change the resolution, e.g., changing the voxel size from 1 mm to 2 mm
Anatomical data, for example an anatomical MRI or other volumentric representations (such as a segmentation) are represented as a volume MATLAB structure in FieldTrip.
An example volume structure is
% read in the tutorial MRI data
mri = ft_read_mri('Subject01.mri');
disp(mri)
dim: [256 256 256] % the size of the 3D volume in voxels
anatomy: [256x256x256 int16] % the numeric data, in this case anatomical information
transform: [4x4 double] % affine transformation matrix
coordsys: 'ctf' % description of the (head) coordinate system
hdr: [1x1 struct]
This volume is already aligned to the CTF head coordinate system. But the alignment by itself does not change the orientation of the original anatomy field, it just adds a transformation matrix which allows for relating each voxel intensity to the corresponding voxel location in the CTF head coordinate system. Note that the transformation matrix is taken into account at all subsequent computations.
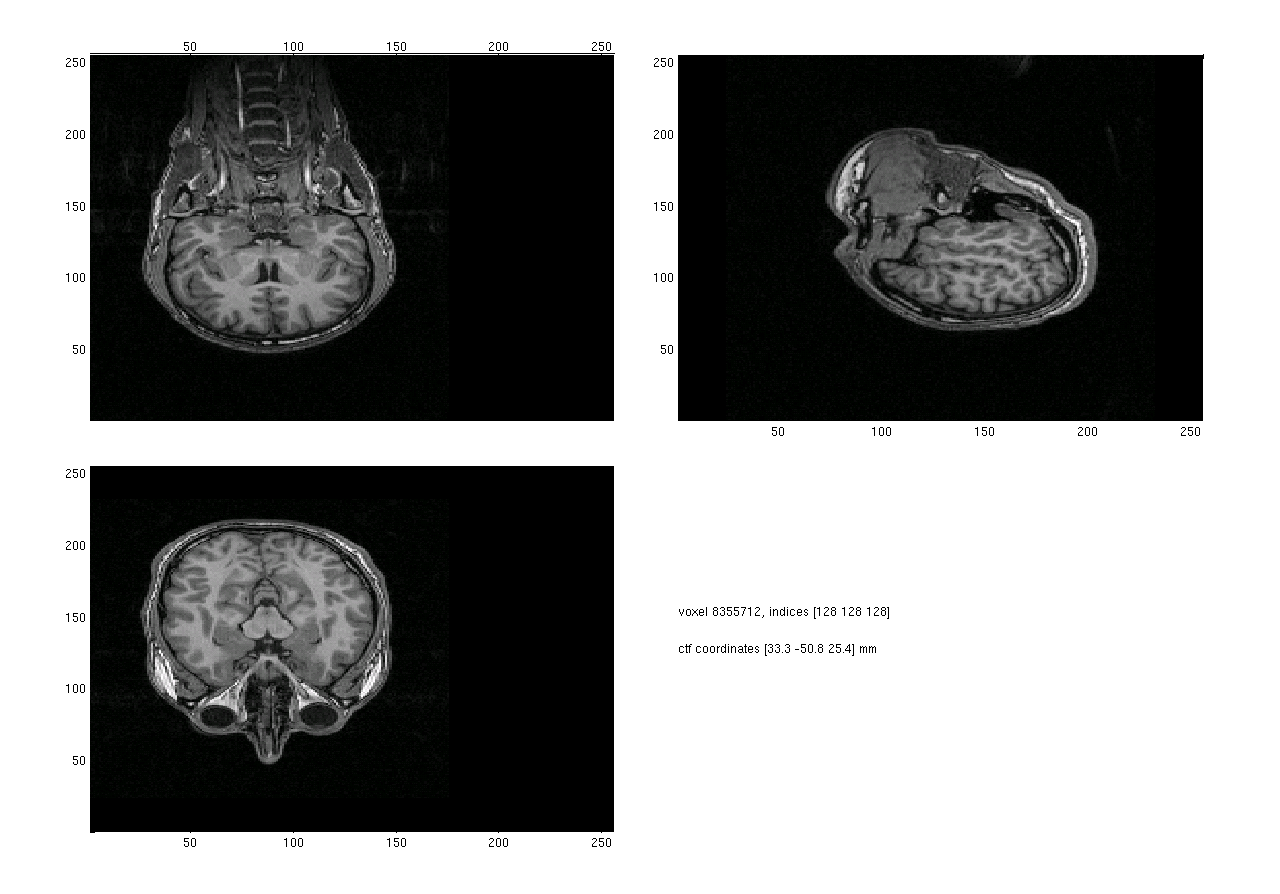
When the anatomical data is plotted, the default behavior is to plot the anatomical data as it is in the anatomy field, without changing its orientation according to the transformation matrix. This is the reason why anatomical data is sometimes plotted with the top of the head pointing to the bottom of the screen (figure 1).
cfg = [];
ft_sourceplot(cfg,mri)
Figure 1. Anatomical mri plotted without using ft_volumereslice before plotting
When you call the ft_volumereslice function on the anatomical MRI, it will apply the transformation matrix to the field anatomy and interpolate the anatomical data onto a new voxel-grid that is aligned with the axis of the head coordinate system. If the input MRI has a coordsys field, the center of the volume will be shifted (with respect to the origin of the coordinate system) for the brain to fit nicely in the box. By default the voxel resolution is 1 mm. The output will have a different orientation of the anatomy and consequently, a different transformation matrix.
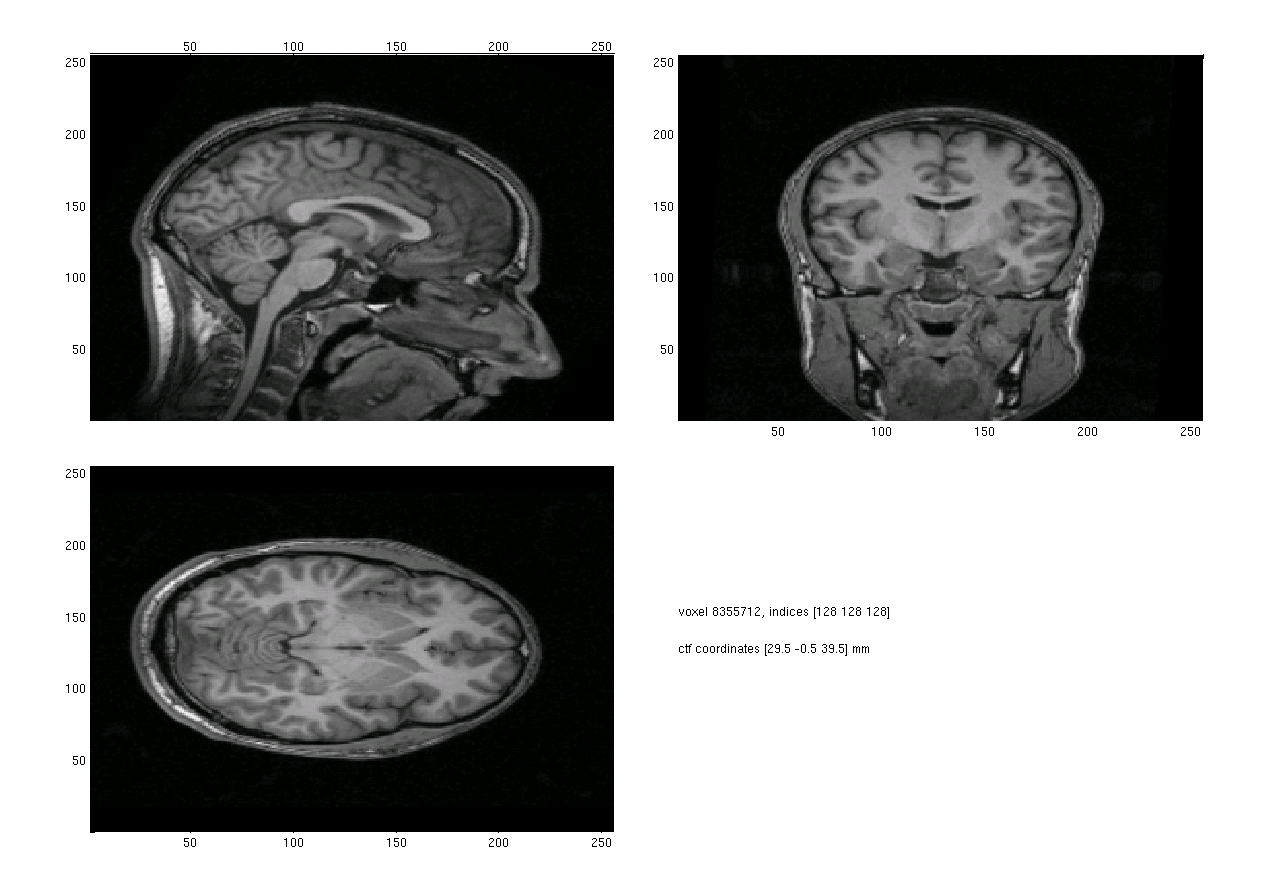
Plotting the resliced anatomical MRI results in a figure with the usually desired orientation of the head (Figure 2).
cfg = [];
mrirs = ft_volumereslice(cfg,mri);
cfg = [];
ft_sourceplot(cfg,mrirs);
Figure 2. Plot of the anatomical mri after using ft_volumereslice
Make the voxels isotropic
The ft_volumereslice function also ensures that the voxels are isotropic (i.e. of equal distance in all three directions). Voxels of an MRI scan are not isotropic when there is a different voxel resolution within the MRI slices than the gap size between the slices. This means that the size of a voxel is not equal in all three directions.
Isotropic voxels are necessary if we want to apply morphological operators to the anatomical volume (e.g., dilating, opening, etc.). These operations occur for example when the skull tissue is segmented in an anatomical volume (e.g., see this tutorial).
Change the field-of-view (FOV)
The ft_volumereslice function is also able to change the number of voxels along each direction. This can be useful for example, when the preprocessing of the anatomical images requires a specific image size (e.g., see this tutorial).
In the figures above you can appreciate the change in the FOV by considering the MRI in the original representation not being in the centre of the picture, whereas after reslicing it is in the centre and better fills the available space.
General principle of reslicing as a 3D interpolation
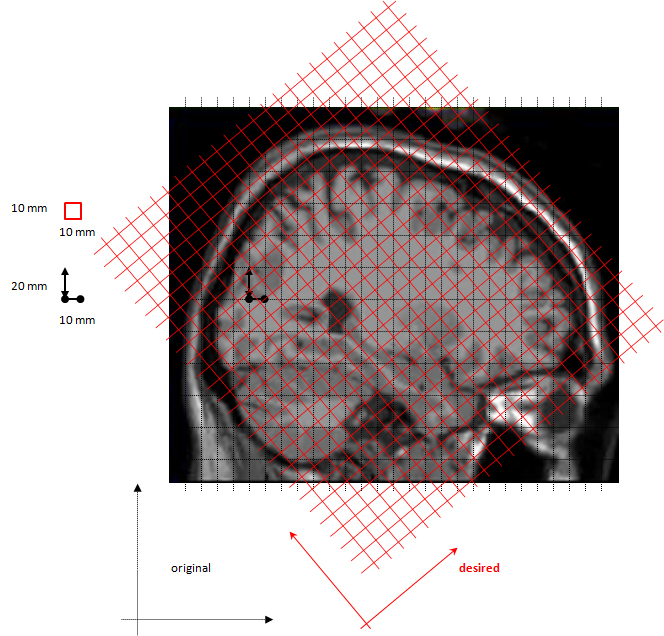
The ft_volumereslice operations, such as changing the orientation of the anatomy and changing the resolution of the voxels can be conceptually understood by looking at the figure below:
The figure shows the original volumetric slices (dotted black lines) and the desired slices (bold red). Note that the distance between the original slices is 2 cm, whereas the pixel distance within the same slice is 1 cm (black). After re-slicing (red) the voxels’ dimensions are the same. You can also see that the voxels are aligned with the axes of the coordinate system to which the image was re-aligned earlier (see the black vs. red axes at the bottom of the figures).