workshop / madrid2019 / tutorial_cleaning /
Cleaning and processing resting-state EEG
Introduction
This tutorial shows how to preprocess and analyze resting state EEG data using an open access resting state EEG dataset that is shared by the University of Cambridge. You can click here for details on the dataset. In this tutorial you will learn how to load and inspect this dataset using FieldTrip. You will perform some basic preprocessing such as repairing broken channels, visual artifact rejection and artifact correction using ICA.
Background
We will adapt the pipeline described in de Cheveigne & Arzounian (2018) Robust detrending, rereferencing, outlier detection, and inpainting for multichannel data. They discuss different algorithms to preprocess MEG or EEG data and - importantly - they propose rules of thumb regarding the order on which these preprocessing steps should be applied. Please, bear in mind that the present pipeline is quite general and, as such, may not apply to specific cases. Please read de Cheveigne & Arzounian (2018) thoroughly and follow this tutorial with a critical mind.
Let us begin with the rules of thumb proposed by de Cheveigne and Arzounian:
As a rule of thumb, if algorithm B is sensitive to an artifact that algorithm A can remove, then A should be applied before B. A difficulty arises of course if A is also sensitive to artifacts that B can remove.
A likely sequence might be:
- discard pathological channels for which there is no useful signal,
- apply robust detrending to each channel,
- detect and interpolate temporally-local channel-specific glitches,
- robust rereference,
- project out eye artifacts (e.g., using ICA or DSS),
- fit and remove, or project out, 50 Hz and harmonics,
- project out alpha activity, etc.,
- apply linear analysis techniques (ICA, CSP, etc.) to further isolate activity of interest.
With these guidelines in mind, let us take the Chennu et al., dataset and begin with the cleaning.
Procedure
In this tutorial the following steps will be taken:
- Read the data into MATLAB using ft_preprocessing and visualize the data in between processsing steps with ft_databrowser
- Interpolate broken channels or noisy data segments with ft_channelrepair, removing artifacts with ft_rejectartifact
- Select relevant segments of data using ft_redefinetrial as well as concatenating data using ft_appenddata
- Once all data is cleaned, correct for eye movement artifacts by running independent component analysis using ft_componentanalysis
Reading in data
The dataset that has been shared does not consist of the original recordings; the data has been imported and some preprocessing steps have been performed already (EEG channels were demeaned and band-pass filtered between 0.5 Hz - 45 Hz, some channels were interpolated using spherical spline algorithms and the data were rereferenced to the average taken over all channels; see Materials and Methods’ section).
For this tutorial we will use the EEG data from one example subject (subj22), which has been selected because the data still shows some artifacts. You can download both raw and processed data of the example subject here.
If you are interested in the raw data from all subjects transformed into BIDS format, you can download it from our download server. Please note that you do not have to download all subjects for this tutorial.
Our goal now is to identify these noisy periods, eye movements, blinks, muscular artifacts and any other channel-specific abnormal behavior. We will guide you through the preprocessing pipeline with the data of the example subject and for one experimental block, i.e., level of sedation:
subj = 'sub-22';
We will start with some minimal preprocessing:
cfg = [];
cfg.dataset = ['/madrid2019/tutorial_cleaning/single_subject_resting/' subj '_task-rest_run-3_eeg.vhdr'];
cfg.channel = 'all';
cfg.demean = 'yes';
cfg.detrend = 'no';
cfg.reref = 'yes';
cfg.refchannel = 'all';
cfg.refmethod = 'avg';
data = ft_preprocessing(cfg);
As you notice we use the average reference, which is breaking with de Cheveigne & Arzounian’s guidelines. We do it anyways because the average reference enhances the interpretability of the data when using ft_databrowser. For example, interpreting topoplots to identify both artifacts and interesting electrophysiological phenomena is always easier with an average reference. Keep in mind though that by subtracting an average, noisy periods present in only few EEG channels will spread to all other channels.
Following preprocessing, the data will have the following fields
data =
hdr: [1x1 struct]
label: {91x1 cell}
time: {[1x90000 double]}
trial: {[91x90000 double]}
fsample: 250
sampleinfo: [1 90000]
cfg: [1x1 struct]
Subsequently we will add the electrode description. For this we will use a custom script which is included with the data on the download server. The main reason for a custom script is that the EEG cap used in this study is the Electrical Geodesics Inc. (EGI) geodesic net, which has its own specific nomenclature for electrode positions, but some of the electrode positions correspond to the 10-10 standard system. If you are curious about the equivalence between the two systems, take a look at the custom function and here.
data.elec = prepare_elec_chennu2016(data.label);
Discarding pathological channels for which there is no useful signal is the first step in de Cheveigne & Arzounian’s guidelines, so we will now use the ft_databrowser to visually inspect our data and mark time segments where data is noisy. We can use this function to simultaneously mark and keep track of different types of physiological artifacts, such as blinks or muscle artifacts. If you have not used the databrowser before, read here how to use it.
cfg = [];
cfg.channel = 'all';
cfg.layout = ft_prepare_layout([],data);
cfg.viewmode = 'vertical';
cfg.blocksize = 5; % time window to browse
cfg.artifactalpha = 0.8; % this make the colors less transparent and thus more vibrant
cfg.artfctdef.blinks.artifact = [];
cfg.artfctdef.badchannel.artifact = [];
cfg.artfctdef.muscle.artifact = [];
artif = ft_databrowser(cfg, data);
Exercise 1
Browse through the segments to get a feel for the data. Do you see any obvious artifacts? There is one channel carrying several artifacts throughout the recording, can you find it? Use the identify button to see the channel name.
We manually add the names of channels that are bad throughout the entire recording to the artifact structure. Many FieldTrip functions, ie ft_channelselection or ft_channelrepair, which we will use further _down, require the specification of channel names in their configuration. For _this specify channel names as strings in a cell-array such as {‘E7’;’Oz’}
artif.badchannel = input('write badchannels: ');
artif.misschannel = input('write missed channels: ');
to save disk space and to prevent doing the same interactive work twice, it is advisable to save the minimal information and run the pipeline again to reconstruct the data. Here, we have already saved the artifacts we identified but not the cleaned EEG data. You can either load the preselected artifact file ‘sub-22_run-03_eeg_artif’ or continue with your own selection.
load('/madrid2019/tutorial_cleaning/sub-22_run-03_eeg_artif')
Interpolating bad channels
We will now explore two different ways of dealing with noisy channels. One is to interpolate entire channels. The other way is to interpolate only segments that contain artifacts.
We repair channels by interpolation, or more precisely by replacing the bad channel with the average of their neighbouring channels. This requires a definition of neighbouring channels. FieldTrip comes with a variety of templates for defining neighbouring channels. For this dataset we provide this information for you, but in general you know your own EEG system and your own data best and you should therefore think about your own neighbours structure. See also the ft_prepare_neighbours function.
load('/madrid2019/tutorial_cleaning/cfg_neighbours', 'neighbours');
Interpolate channels that are bad during the whole experiment
cfg = [];
cfg.badchannel = artif.badchannel;
cfg.method = 'weighted';
cfg.neighbours = neighbours;
data_fixed = ft_channelrepair(cfg,data);
For this subject the noisy channel only has a handful of artifacts, so instead of interpolating an entire channel, we will only interpolate the noisy segments.
Interpolate bad channels for specific segments
Following de Cheveigne and Arzounian’s third step, we are going to detect and interpolate temporally-local channel-specific glitches.
Note we deliberately skip step 2. apply robust detrending to each channel for later because it is necessary to find first the pieces of data with artifacts and to exclude them. De Cheveigne and Arzounian’s detrending algorithm has the possibility to exclude outliers so this is the main reason for us to change the order (check their nt_detrend.m function).
We can make a selection of the segments in which one of the channels was bad.
artpadding = 0.1;
begart = artif.artfctdef.badchannel.artifact(:,1)-round(artpadding.*data.fsample);
endart = artif.artfctdef.badchannel.artifact(:,2)+round(artpadding.*data.fsample);
offset = zeros(size(endart));
% do not go before the start of the recording or the end
begart(begart<1) = 1;
endart(endart>max(data.sampleinfo(:,2))) = max(data.sampleinfo(:,2));
cfg = [];
cfg.trl = [begart endart offset];
data_bad = ft_redefinetrial(cfg, data);
You should now have a data structure data_bad, that contains the segments of data you have identified as artifact in the trial field. Note that each bad segment and hence each trial will be of different length. Inspect the fields in your data structure and compare them to the original data structure.
data_bad =
hdr: [1x1 struct]
trial: {[91x395 double] [91x333 double]}
time: {[1x395 double] [1x333 double]}
elec: [1x1 struct]
fsample: 250
label: {91x1 cell}
sampleinfo: [2x2 double]
cfg: [1x1 struct]
Subsequently we identify the channels with the artifacts using the algorithm by de Cheveigne (see nt_find_bad_channels.m in Noisetools).
The following (rather complicated) piece of code is a mix of regular MATLAB code with FieldTrip functions. If it is useful, we might add it to one of the FieldTrip functions later to make it easier to use.
% The parameters to detect artifacts are:
proportion = 0.4; % criterion proportion of bad samples
thresh1 = 3; % threshold in units of median absolute value over all data
data_fixed = {};
for k=1:size(data_bad.trial,2)
w = ones(size(data_bad.trial{1,k}));
md = median(abs(data_bad.trial{1,k}(:)));
w(find(abs(data_bad.trial{1,k}) > thresh1*md)) = 0;
iBad = find(mean(1-w,2)>proportion);
[val iBad_a] = max(max(abs(data_bad.trial{1,k}.*(1-w)),[],2));
if isempty(iBad)
iBad = find(mean(1-w,2)==max(mean(1-w,2)));
warning(['decreasing threshold to: ' num2str(max(mean(1-w,2)))]);
end
% we use ft_channelrepair to interpolate these short selected artifacts
cfg = [];
cfg.badchannel = data_bad.label([iBad;iBad_a]);
cfg.method = 'weighted';
cfg.neighbours = neighbours;
cfg.trials = k;
data_fixed{1,k} = ft_channelrepair(cfg, data_bad);
end
After correcting each artifactual trial we can use ft_appenddata to combine the trials into one structure again.
data_fixed = ft_appenddata([], data_fixed{:});
Exercise 2
Visualize the selected artifacts in data_bad with
ft_databrowser and compare it to data_fixed.
You can also just plot the bad channel by specifying its name in cfg.channel. Or
you can use the standard MATLAB plot function. For this you need to find the
index of the channel using the data.label field.
Returning to the original data, we now delete the segments that contain artifacts and append the fixed data.
clear data_bad
cfg = [];
cfg.artfctdef.minaccepttim = 0.010;
cfg.artfctdef.reject = 'partial';
cfg.artfctdef.badchannel.artifact = [begart endart];
data_rejected = ft_rejectartifact(cfg, data);
data = ft_appenddata([], data_rejected, data_fixed);
% clear these variables from memory to avoid confusion later on
clear data_rejected data_fixed
Exercise 3
Inspect the new data structure. What has changed?
In order keep the data as one continuous trial we use ft_redefinetrial and the sample information
cfg = [];
cfg.trl = [min(data.sampleinfo(:,1)) max(data.sampleinfo(:,2)) 0];
data = ft_redefinetrial(cfg,data);
Visualize the results of channel interpolation
We can use ft_databrowser to check the results of the interpolation.
cfg = [];
cfg.viewmode = 'vertical';
cfg.artifactalpha = 0.8;
cfg.blocksize = 5;
if isfield(artif.artfctdef,'badchannel')
cfg.artfctdef.badchannel.artifact = artif.artfctdef.badchannel.artifact;
end
if isfield(artif.artfctdef,'visual')
cfg.artfctdef.visual.artifact = artif.artfctdef.visual.artifact;
end
if isfield(artif.artfctdef,'muscle')
cfg.artfctdef.muscle.artifact = artif.artfctdef.muscle.artifact;
end
ft_databrowser(cfg,data);
Reject the muscular and visual artifacts
Eye movements and blinks cause artifacts in the EEG data because the retina (which is electrically charged) moves when the subject blinks. The contribution of eye artifacts to the EEG data can either be detected manually and cut out or be estimated and removed with independent component analysis (ICA).
If you have marked muscular or visual artifacts manually, you can reject (i.e. cut out and remove) the noisy segments like this:
cfg = [];
cfg.artfctdef.minaccepttim = 0.010;
cfg.artfctdef.reject = 'partial';
if isfield(artif.artfctdef,'visual')
cfg.artfctdef.visual.artifact = artif.artfctdef.visual.artifact;
end
if isfield(artif.artfctdef,'muscle')
cfg.artfctdef.muscle.artifact = artif.artfctdef.muscle.artifact;
end
data = ft_rejectartifact(cfg, data);
Detrending the dataset
At this point, given that we already excluded or interpolated the pieces of EEG data containing artifacts, it is safer to detrend the data. As mentioned above, the robust detrending de Cheveigne and Arzounian proposed in their paper is a single function (nt_detrend.m) that detects outliers and do not take them into account during the detrend operation. In FieldTrip we do not have a dedicated function to do this but we have developed this pipeline (see above) to exclude artifacts and be able to perform the detrending as follows:
cfg = [];
cfg.channel = 'all';
cfg.demean = 'yes';
cfg.polyremoval = 'yes';
cfg.polyorder = 1; % with cfg.polyorder = 1 is equivalent to cfg.detrend = 'yes'
data = ft_preprocessing(cfg, data);
So now we performed the second step of the guidelines: 2. apply robust
detrending to each channel,. You can play with the cfg.polyorder parameter and
check what happens with the data.
Robust rereference
All the data has been cleaned and now we can archive a more robust rereference, so step 4 can be done as follows:
cfg = [];
cfg.channel = 'all';
cfg.demean = 'yes';
cfg.reref = 'yes';
cfg.refchannel = 'all';
cfg.refmethod = 'avg';
data = ft_preprocessing(cfg, data);
Eye artifact removal with ICA
This brings us to step 5. project out eye artifacts (e.g., using ICA or DSS). A reliable ICA decomposition requires as much data as possible. In theory, the spatial distribution of the eye artifacts should not be different in the different experimental conditions. However, the estimate of the independent components (IC) can be unstable and the more data is available the more reliable they become. Importantly, the more channels you record the more time-points you will need for reliable estimates (for empirical data on this issue, see Groppe et al., 2009. Therefore to improve the estimate and to make sure that you are removing the same eye activity in all conditions, you should combine data from the different runs/conditions.
We concatenate all trials into one matrix to compute the rank of the data, this helps to constrain the number of independent components.
dat = cat(2, data.trial{:});
dat(isnan(dat)) = 0;
n_ic = rank(dat);
% Groppe et al. reliability estimate
size(dat,2)/(91^2)
cfg = [];
cfg.method = 'runica';
cfg.numcomponent = n_ic;
comp = ft_componentanalysis(cfg, data);
Because computing the IC can be time consuming, it is efficient to save the result. To reduce disk space we delete the ‘time’ and ‘trial’ fields, because with the ‘topo’ and ‘unmixing’ matrix we can reconstruct everything.
comp = rmfield(comp, 'time');
comp = rmfield(comp, 'trial');
save([subj 'run-03_comp.mat'], '-struct', 'comp');
In general we recommend to not change the FieldTrip structures. It increases the chances of accidental data corruption and errors later in your analysis pipeline.
You can load the pre-computed topo and unmixing matrix if you do not want to wait for the ICA to finish. Those have been computed on all four runs (sedation levels) combined. Using the pre-computed topo and unmixing matrix, we can quickly redo the ICA unmixing.
comp = load(['/madrid2019/tutorial_cleaning/,subj,'_comp']);
cfg = [];
cfg.demean = 'no'; % This has to be explicitly stated, as the default is to demean.
cfg.unmixing = comp.unmixing; % Supply the matrix necessary to 'unmix' the channel-series data into components
cfg.topolabel = comp.topolabel; % Supply the original channel label information
comp = ft_componentanalysis(cfg, data);
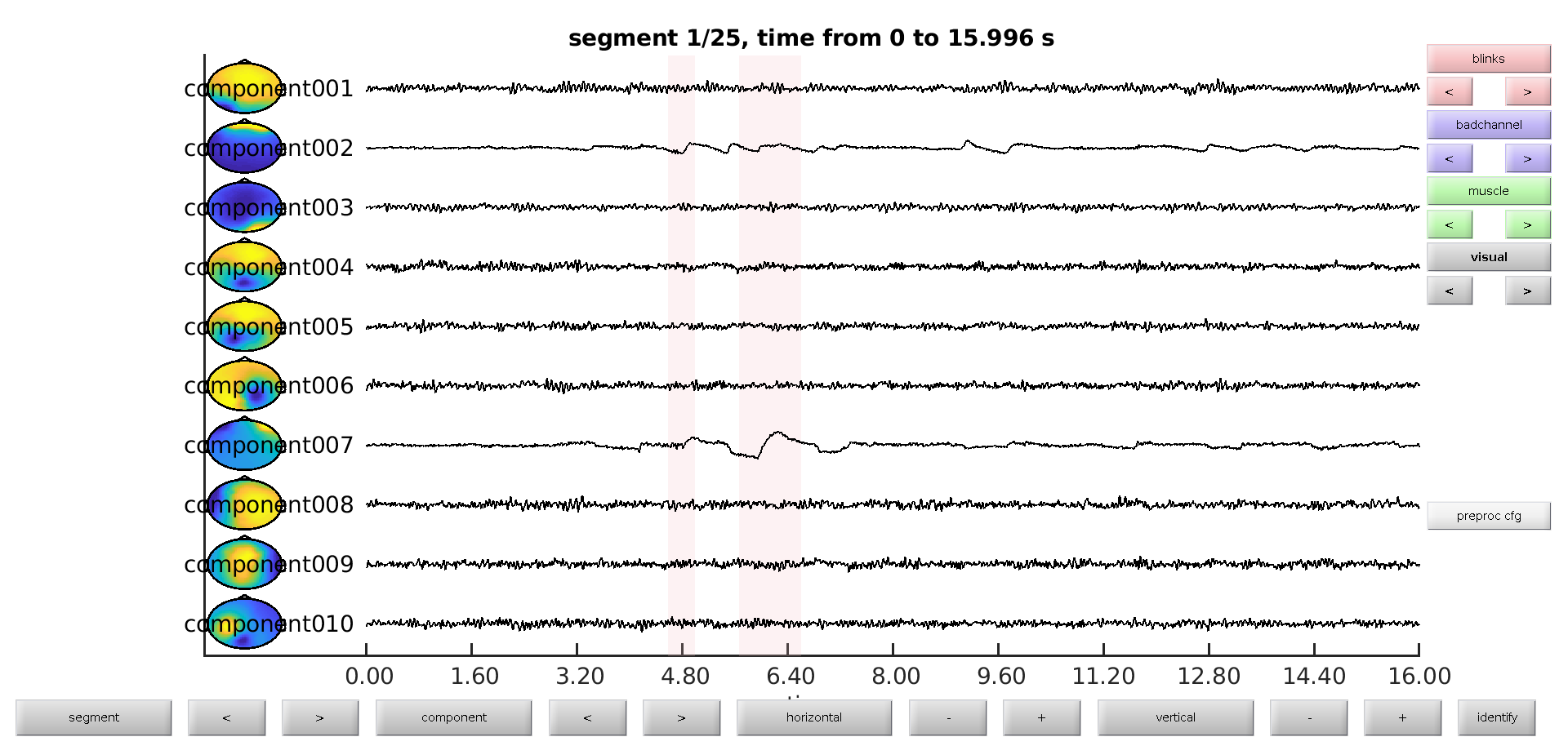
We now visualize the components to select which ones model the artifacts. We add the visually identified artifacts in order to see, which of those are matched by the independent component’s time course.
data.elec = prepare_elec_chennu2016(data.label);
cfg = [];
cfg.layout = ft_prepare_layout([],data);
cfg.viewmode = 'component';
cfg.zlim = 'maxmin';
cfg.compscale = 'local'; % scale each component separately
cfg.contournum = 6;
cfg.artifactalpha = 0.8;
cfg.artfctdef = artif.artfctdef;
ft_databrowser(cfg, comp);
ic.selected = input('ICs to keep (i.e. [1 5]): ');
ic.artifact = input('ICs to reject (i.e. [8]): ');
save([subj '_ic_selection.mat'],'-struct','ic');
cfg = [];
cfg.component = ic.artifact;
data = ft_rejectcomponent(cfg, comp, data);
Exercise 4
Use ft_databrowser one last time to view the cleaned data. Did the ICA successfully correct all eye blinks?
Remove 50 Hz line noise
Step 6. fit and remove, or project out, 50 Hz and harmonics, does not make sense for this dataset, because it was already bandpass filtered (0.5 Hz - 45 Hz). In case you are interested, the line noise cleaning can be perform by fitting sine waves (cfg.dftfilter) at specific frequencies specified in cfg.dftfreq or by using a band stop filter (cfg.bpfilter):
cfg = [];
cfg.channel = 'all';
cfg.demean = 'yes';
cfg.dftfilter = 'yes';
cfg.dftfreq = [50];
% cfg.bsfilter = 'no'; % band-stop method
% cfg.bsfreq = [48 52];
data = ft_preprocessing(cfg,data);
See also
A more general overview of dealing with artifacts is provided in the artifact tutorial.